My research is in the area computational biology and bioinformatics, parallel and distributed high performance computing. I am particularly interested in network models of protein interactions, and their application in understanding diseases and drug design. In the area of parallel programming, my work concentrates on parallel programming patterns, teaching techniques for programming multi-core computers, design and development of high performance parallel/distributed algorithms for computational biology.
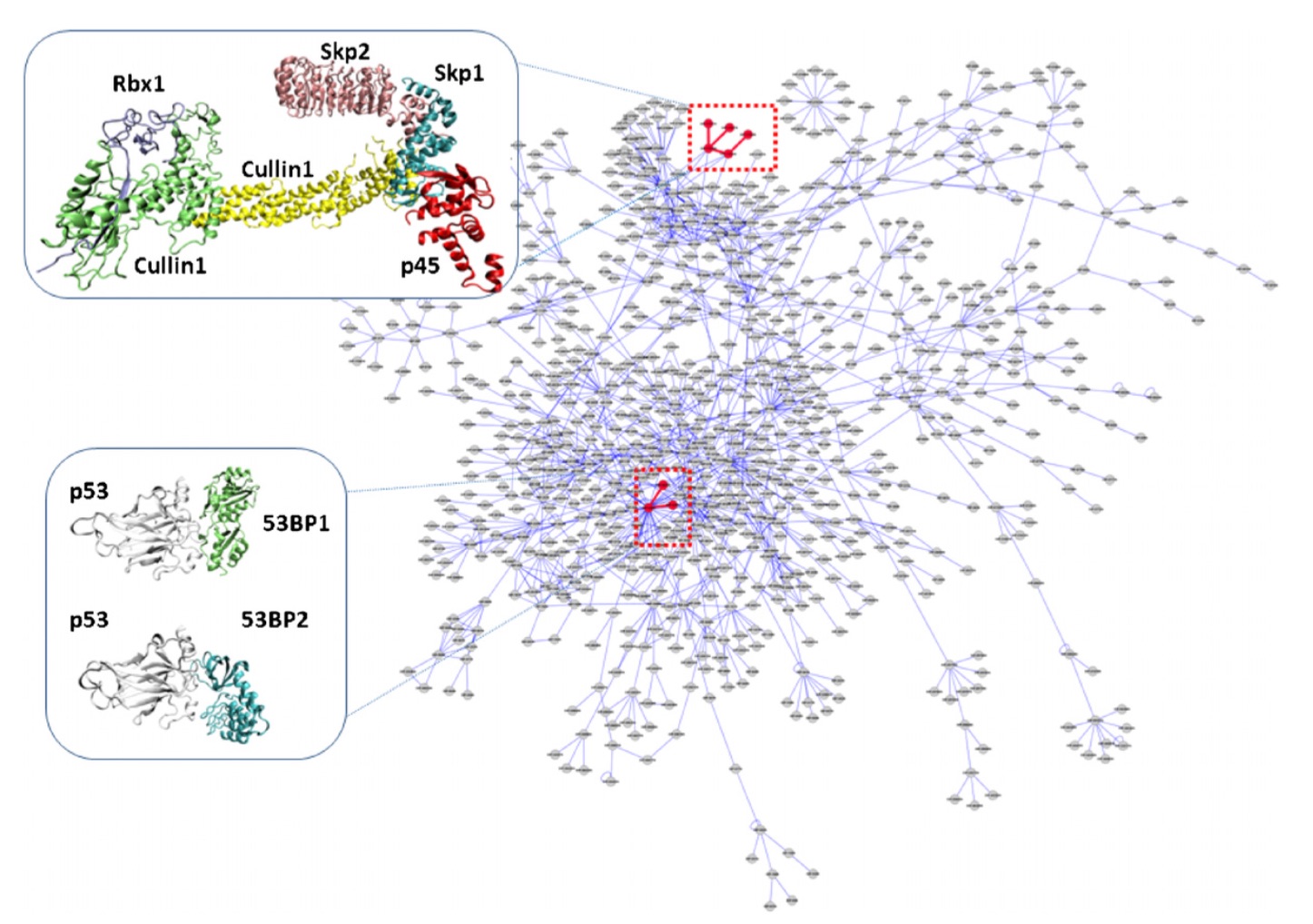
Biology is becoming an information driven science. An explosion of biological data including genome sequences and protein structures has been taking place with the recent advances in biotechnology, hence computational methods have become increasingly important to extract knowledge from complex data. In collaboration with faculty from computational and biological sciences, I am investigating how proteins interact with each other (PRISM project). Proteins are the working horse of the cellular machinery. They are responsible for diverse functions ranging from molecular motors to signaling. Prediction of protein-protein interactions is crucial for understanding complex biological processes as well as understanding diseases and for drug discovery. By unifying protein interfaces with protein-interaction networks, PRISM project aims to provide insight into the role of proteins within complex network of interactions. The research integrates various approaches/methods from machine learning, computer graphics, graph theory, and parallel computing in developing new prediction and analysis methods.
For more information on the research project, please visit our research group COSBI.