RESEARCH
Every cell comes from a cell. That is, in order to obtain a new cell, a pre-existing cell must duplicate its contents and split into two. Indeed, this is simply how all living cells reproduce. They grow, replicate their chromosomes and segregate them to the daughter cells. Finally, the two daughter cells separate by abscission. The series of events taking part in this cell reproduction process is known as the cell cycle (Figure 1), which is may be the most fascinating incident in the fantastic world of the cell. The beauty of the cell cycle lies on elaborate orchestration of discrete events, which requires precise control mechanisms for accuracy and coordination. For the last half-century many studies have been made to understand the basics of the cell division cycle. Yet, we lack the comprehensive view of it.
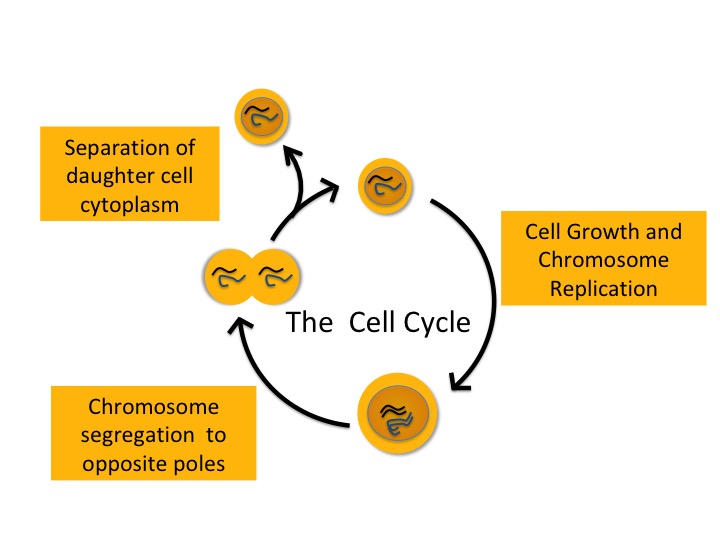
Figure 1. The Cell Cycle. Cells replicate their chromosomes, segregate them to the daughter cells and finally, the two daughter cells separate.
Our research is focused at last stages of the cell cycle. One particular question we ask is how mitotic cell cycle is ended. Eukaryotic cells divide through mitosis to produce new cells with identical DNA. Yet, mitosis has to be ended for the two cells to be separated to become individual cells with the capacity to further divide. This occurs by a process called mitotic exit. Failure to control mitotic exit is the key cause of aneuploidy and genomic instability, which are hallmarks of cancer. Our lab is interested in understanding how cells control exit from mitosis to maintain genome integrity. We use the Saccharomyces cerevisiae model in our research.
Saccharomyces cerevisiae as a model organism in cell cycle studies
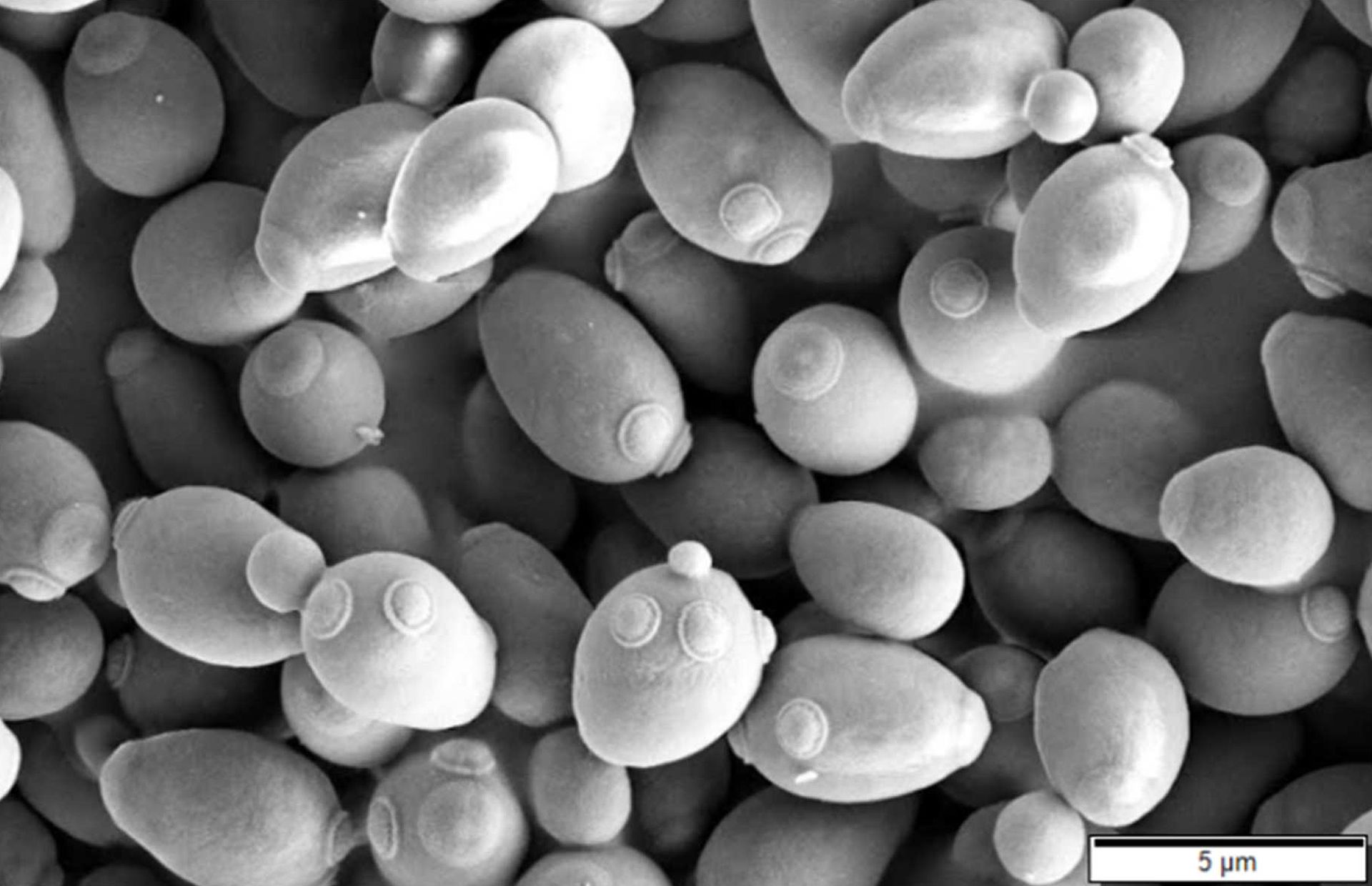

Figure 2. SEM images of budding yeast cells. (https://commons.wikimedia.org/wiki/Category:Microscopic_images_of_Saccharomyces_cerevisiae#/media/File:Saccharomyces_cerevisiae_SEM.jpg)
There are mainly two yeast species widely used as model organisms in cell cycle studies: Schizosaccharomyces pombe and Saccharomyces cerevisiae. S. cerevisiae, known as baker’s or brewer’s yeast and also called as budding yeast, is an oval-shaped single-celled fungus (Figure 2). S. cerevisiae genome with ~13 million base pairs on 16 chromosomes was the first fully sequenced eukaryotic genome ever 3. Budding yeast life cycle consists of alterations between the haploid and diploid states. This makes the haploid laboratory yeast an excellent system to characterize genes through study of their functions by mutating them, which would be difficult in diploids due to presence of two gene copies. In fact, cyclin dependent kinases, the basic regulators of the cell cycle, were discovered first in budding yeast, through such mutational analysis 4.
In addition, genetic manipulation of yeast by means of knocking-out / knocking-in of genes or by tagging the genes in their own loci is also straight-forward, thanks to the high rate of homologous recombination and well-established methods in budding yeast 5-8.
Another striking feature of budding yeast is that every cell division is inherently asymmetric. A cell division is called asymmetric when the two daughter cells differ in size and/or content. In higher eukaryotes asymmetric cell division is very crucial for cellular differentiation and stem cell maintenance. However in budding yeast, asymmetric cell division per se is the way of reproduction. In every cell cycle, there is an established polarity between the mother cell and the bud. Hence both compartments are different in terms of their contents, as well as their size, during and after cell division. Being a unicellular organism with all the advantages mentioned above, budding yeast constitutes an excellent model for cell cycle and asymmetric cell division.
Cell Cycle in Budding Yeast
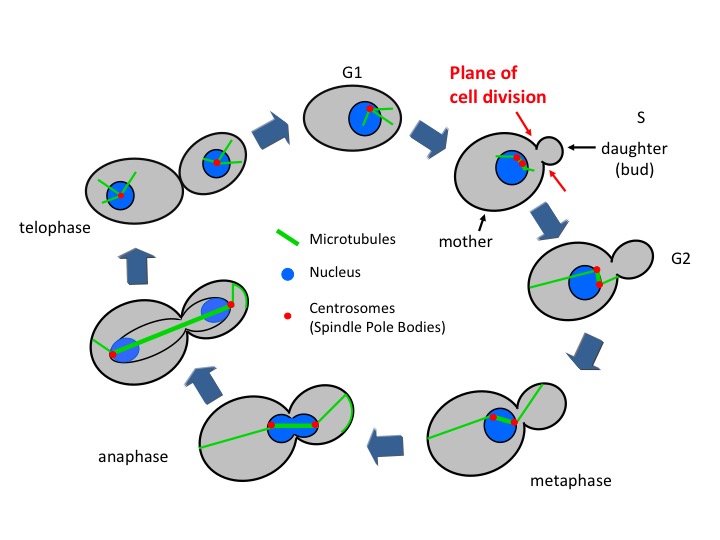
Figure 3. Budding yeast cell cycle
Once yeast cells become committed to cell division, they initiate budding (the processes by which a tiny daughter cell starts appearing and growing on the top of a mother cell, hence the name “budding yeast”). The daughter cell (bud) stays connected with the mother cell until the end of cell cycle. The boundary between them (so called bud neck) also denotes the future site of cytokinesis. Meanwhile yeast cells also duplicate their microtubule organizing center named spindle pole body (SPB, the functional equivalent of the mammalian centrosome) 9 and replicate their chromosomes. Duplicated chromosomes get attached to the microtubules at their kinetochores in a bipolar manner (metaphase) and are pulled apart towards the mother and the daughter compartments so that each cell gets one copy of each chromosome (anaphase). Chromosome segregation is driven by nuclear (spindle) and cytoplasmic microtubules, with the assistance of microtubule binding proteins and molecular motors 10. Unlike higher eukaryotes, yeast undergoes a “closed mitosis” in which the nuclear envelope does not breakdown during mitosis 11. At the end of mitosis, the two cells get physically separated (cytokinesis) by contraction of the actomyosin ring at the bud neck and deposition of membrane and cell wall material at the division site 12-14.
Mitosis, driven by mitotic cyclin/Cdks, requires inactivation of the mitotic cyclin/Cdks and reversal of the Cdk dependent phosphorylation of several Cdk substrates in order to finish. Cdc14 in budding yeast is a dual specificity protein phosphatase that is capable of doing both 15. At anaphase onset, a transient wave of Cdc14 activation is triggered by the cdc Fourteen Early Anaphase Release (FEAR) network; whereas later in anaphase a conserved signaling pathway named the mitotic exit network (MEN) promotes fully activation of Cdc14 that is only reversed by transit into the next cycle. The MEN is essential for mitotic exit, although FEAR is not. A checkpoint called the Spindle Position Checkpoint (SPOC) inhibits the MEN upon spindle mis-orientation, to ensure that chromosome segregation is only triggered once the correct orientation is established. Mitotic exit is also under the control of daughter cell localized factors, albeit in a less defined manner.
Our research aims to identify and characterize novel mechanisms of mitotic exit control.
REFERENCES
- Osborn, M.J. & Miller, J.R. Rescuing yeast mutants with human genes. Brief Funct Genomic Proteomic 6, 104-111 (2007).
- Lee, M.G. & Nurse, P. Complementation used to clone a human homologue of the fission yeast cell cycle control gene cdc2. Nature 327, 31-35 (1987).
- Mewes, H.W. et al. Overview of the yeast genome. Nature 387, 7-65 (1997).
- Hartwell, L.H., Culotti, J., Pringle, J.R. & Reid, B.J. Genetic control of the cell division cycle in yeast. Science 183, 46-51 (1974).
- Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947-962 (2004).
- Knop, M. et al. Epitope tagging of yeast genes using a PCR-based strategy: more tags and improved practical routines. Yeast 15, 963-972 (1999).
- Amberg, D.C., Burke, D. & Strathern, J.N. Methods in yeast genetics : a Cold Spring Harbor Laboratory course manual, Edn. 2005. (2005).
- Longtine, M.S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953-961 (1998).
- Forsburg, S.L. & Nurse, P. Cell cycle regulation in the yeasts Saccharomyces cerevisiae and Schizosaccharomyces pombe. Annu Rev Cell Biol 7, 227-256 (1991).
- Sharp, D.J., Rogers, G.C. & Scholey, J.M. Microtubule motors in mitosis. Nature 407, 41-47 (2000).
- Heath, I.B. Variant mitoses in lower eukaryotes: indicators of the evolution of mitosis. Int Rev Cytol 64, 1-80 (1980).
- Pollard, T.D. Mechanics of cytokinesis in eukaryotes. Curr Opin Cell Biol 22, 50-56 (2010).
- Barr, F.A. & Gruneberg, U. Cytokinesis: placing and making the final cut. Cell 131, 847-860 (2007).
- Balasubramanian, M.K., Bi, E. & Glotzer, M. Comparative analysis of cytokinesis in budding yeast, fission yeast and animal cells. Curr Biol 14, R806-818 (2004).
- Stegmeier, F. & Amon, A. Closing mitosis: the functions of the Cdc14 phosphatase and its regulation. Annu Rev Genet 38, 203-232 (2004).